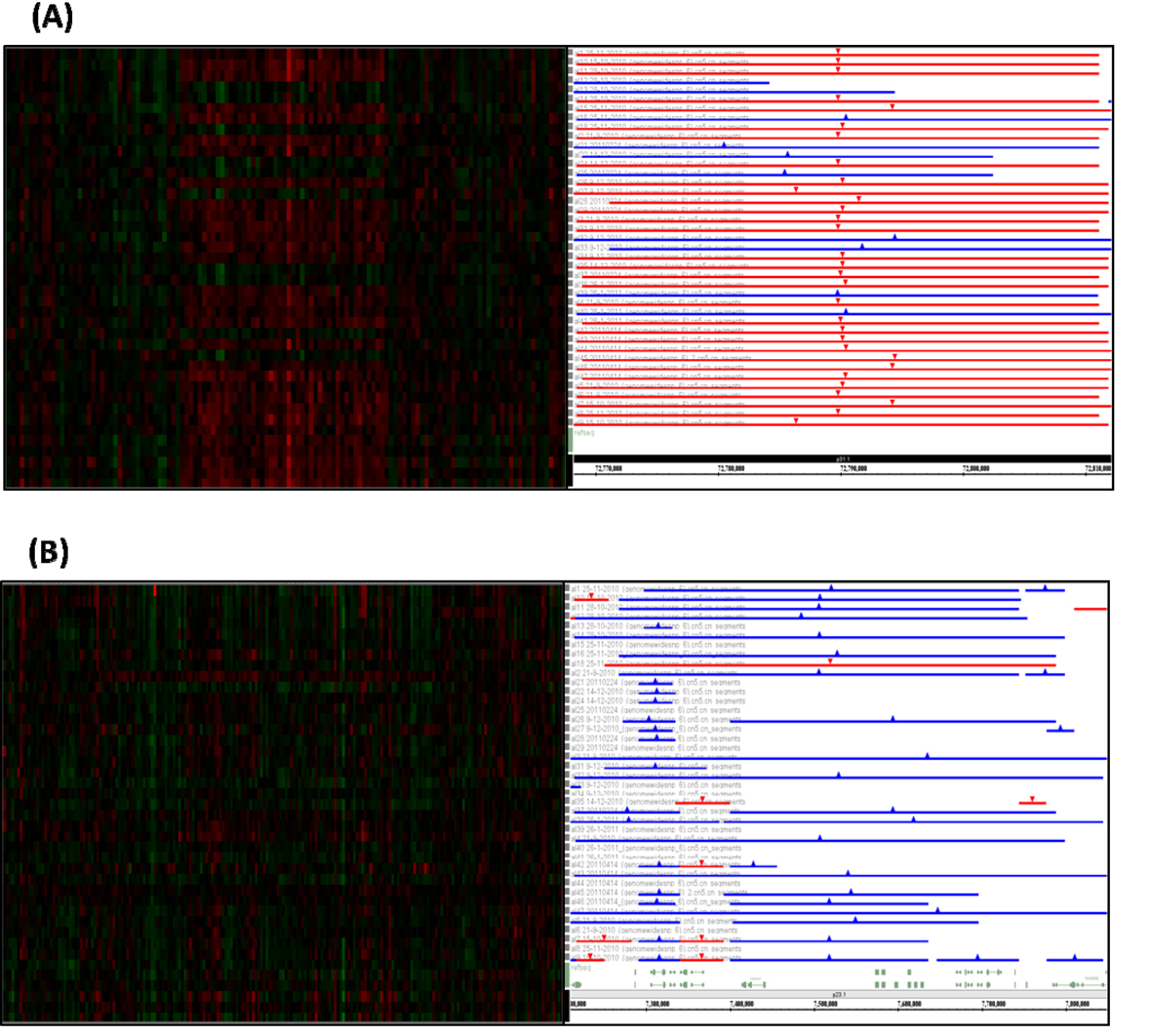
Figure 1. The most common chromosomal alterations in the 41 adult ALL cases. The left lanes are heatmap of Affymetrix SNP array. Red is deletion and green is amplification. The right lanes are segmentation of DNA copy number. Red is deletion and blue is amplification. The most common deletion was found at chromosome chr1p31.1 (A) and amplification was found at chromosome chr8p23.1 (B).
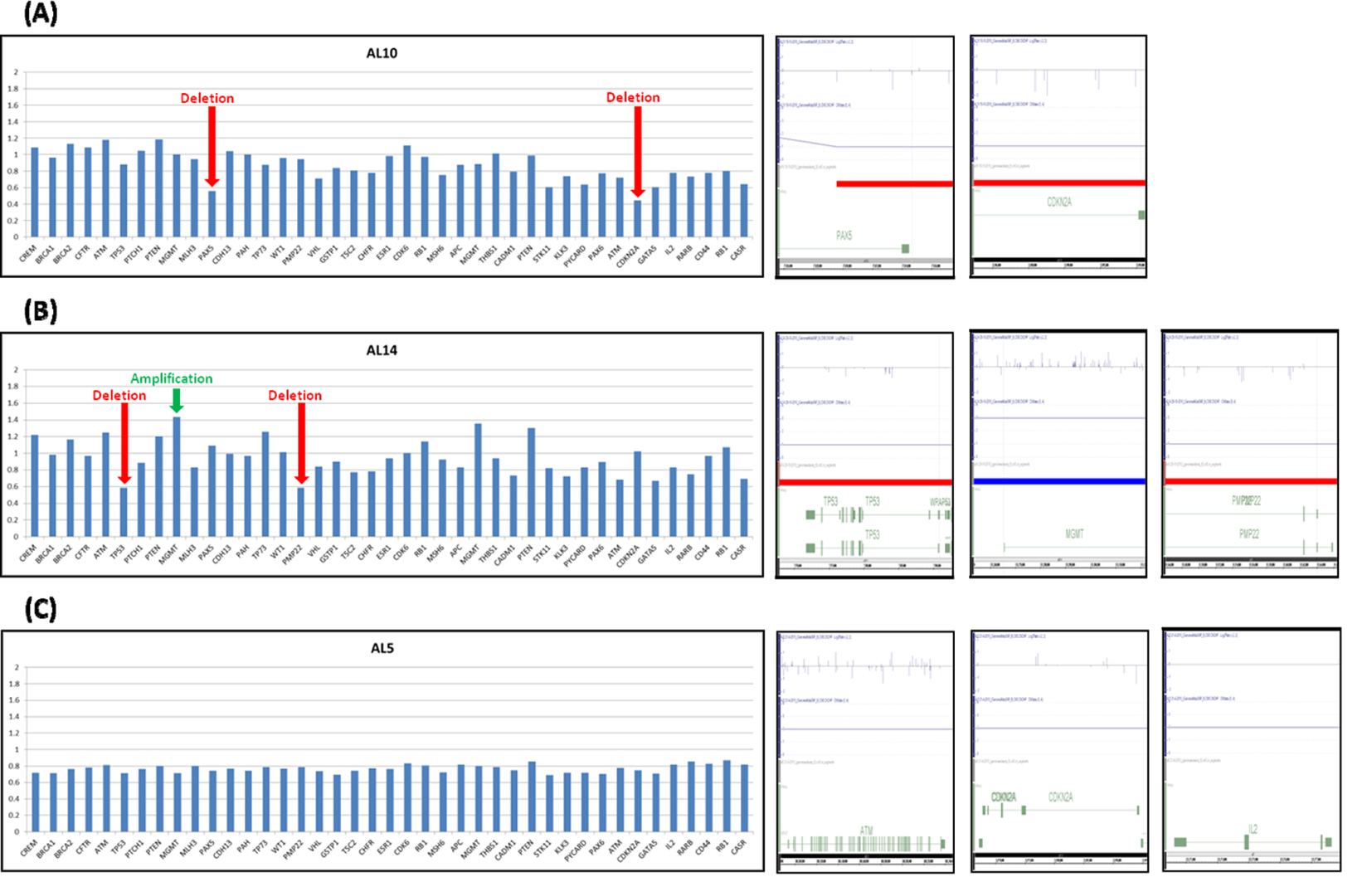
Figure 2. Validation of chromosomal alteration in adult ALL using MLPA technique. The validation results for samples AL10, AL14 and AL5 are labeled as A, B and C respectively. The left lanes showed MLPA results. Ratio < 0.60 was considered as deletion, > 1.40 was considered as amplification. The right lanes showed SNP array results. (A, B) Red arrow indicated that deletion occurred in PAX5 and CDKN2A genes in AL10 sample and TP53 and PMP22 in AL14 sample. (B) Green arrow indicated MGMT gene amplified in AL14 sample. (C) No amplification or deletion was found in AL5 sample by MLPA technique. The results of SNP array were concordance with MLPA result.

