| Journal of Hematology, ISSN 1927-1212 print, 1927-1220 online, Open Access |
| Article copyright, the authors; Journal compilation copyright, J Hematol and Elmer Press Inc |
| Journal website http://www.thejh.org |
Original Article
Volume 6, Number 4, October 2017, pages 73-80
The DAU Allele and Anti-D Alloimmunization Present With High Frequency in Brazilian Sickle Cell Disease Patients
Jose Pereira de Moura Netoa, e, Bruno Antonio Veloso Cerqueirab, Wendell Vilas Boas Santosa, Isa Menezes Lyrad, Marilda Souza Goncalvesc, d
aFaculdade de Farmacia, Universidade Federal do Amazonas, Manaus, Amazonas, Brazil
bDepartamento de Ciencias da Vida, Universidade do Estado da Bahia, Salvador, Bahia, Brazil
cFundacao Oswaldo Cruz - Centro de Pesquisas Goncalo Moniz, Salvador, Bahia, Brazil
dUniversidade Federal da Bahia, Salvador, Bahia, Brazil
eCorresponding Author: Jose Pereira de Moura Neto, Universidade Federal do Amazonas, Faculdade de Ciencias farmaceuticas, Avenida General Rodrigo Otavio Jordao Ramos, 6200 - Coroado I, Manaus - AM, 69067-005, Brazil
Manuscript submitted January 8, 2017, accepted February 14, 2017
Short title: Sickle Cell Disease and DAU Allele
doi: https://doi.org/10.14740/jh316w
| Abstract | ▴Top |
Background: Antigens DIIIa, DAR and DAU are common in people of African descent and are involved in anti-D alloimmunization. Sickle cell disease (SCD) patients frequently need blood therapy and are vulnerable to alloimmunization.
Methods: The study included SCD patients from the Brazilian state of Bahia, which has the highest incidence of the disease in Brazil; 241 SCD patients and 220 healthy individuals were studied. Alleles were characterized by PCR-RFLP and PCR-SSP techniques.
Results: The DAU allele was found in 22.3% (43/193) of the SCD patients. Two (1%) patients had the DIIIa/D wild-type genotype, one (0.5%) had the DIIIa/D- genotype, 11 (5.7%) had the DAR/D wild-type genotype and three (1.6%) had the DAR/D- genotype. Two patients were positive for the 667T>G mutation and the 1136C>T mutation, one (0.5%) had the genotype DIIIa/DAU, and one (0.5%) had the genotype DAR/DAU.
Conclusion: There was statistical significance when the allele frequencies were evaluated among SCD, sickle cell anemia (HbSS) patients and healthy individuals. The frequencies of the DIIIa, DAR and DAU alleles among SCD patients differ from those of healthy individuals from the same population, and a high frequency of the DAU variant was associated with anti-D alloimmunization in these patients.
Keywords: Sickle cell disease; Anti-D alloimmunization; DIIIa; DAR; DAU alleles
| Introduction | ▴Top |
The Rh system is polymorphic and is composed of antigens characterized by high immunogenicity, especially the D antigen. The D antigen (ISBT 004001; RH1) is a red blood cell (RBC) membrane antigen associated with a blood group that is of great importance in blood therapy, because Rh-negative individuals easily develop antibodies when exposed to D-positive RBC [1].
The partial D phenotype is characterized by the absence of one or more epitopes of the original D antigen that were replaced by other amino acid sequences. Those changes are promoted mostly by missense point mutations in the RHD gene or by gene rearrangements between the RHD and RHCE genes due to the high homology and the short distance between them, and they qualitatively change the D protein in its extracellular localization [2-4].
Partial D antigens are rare in the Caucasian population, with frequencies of less than 1%, but are frequent among Africans and individuals of African descent [2-4]. The variant DIIIa was first described in 1996 as one of the six categories of variants of the Rh-positive antigen among individuals who developed anti-D antibody [5]. The DAR variant is characterized by the complete absence of at least nine of the 37 epitopes and is commonly found in Rh-positive individuals. The DAU variant is common in people of African descent, but it is rare in Caucasians. In 1989, du Toit et al conducted a study in the region of Cape Town, South Africa, and reported that 11% of pregnant women were anti-D Rh-positive [6].
Sickle cell disease (SCD) is a genetic disease characterized by an autosomal recessive inheritance. It shows a high incidence and prevalence in Brazil, especially in the Bahia state [7]. Sickle cell anemia (HbSS) is the most severe type of SCD, and individuals with this need frequent blood therapy and are vulnerable to alloimmunization, which elevates the risk of blood transfusion reactions that can be severe or even fatal due to overall organ damage [4].
Therefore, the present study aimed to investigate the frequencies of the DIIIa, DAR and DAU alleles associated with the partial D antigen among SCD patients from the Northeast of Brazil and to estimate the risk of anti-D alloimmunization.
| Materials and Methods | ▴Top |
Study population
A cross-sectional study was developed during the period of 2003 - 2007. The population studied was composed of 241 SCD patients selected from the Hematology Outpatient Clinic of the Fundacao de Hematologia e Hemoterapia do Estado da Bahia (HEMOBA). All patients were phenotyped as D positive or D weak. Clinical data were collected from the medical records, and demographic data were provided by interviews with the patients or with their parents or guardians when the patient was less than 21 years of age. The eligibility criteria included only SCD patients. All patients were at the steady state of the disease, which was characterized as a period without any acute events, and had not had blood therapy prior to blood sampling. The exclusion criteria included the presence of infectious disease, previous blood therapy (less than 4 months before the study) and inflammatory episodes during the study.
The study also included a group of 220 healthy individuals from the same population as the patient group. This group was included to test the frequency of the studied alleles in the same general population. The healthy group was composed of volunteers selected by the Clinical Analysis Laboratory of the Faculdade de Farmacia of Universidade Federal da Bahia (UFBA).
The study was approved by the Centro de Pesquisa Goncalo Moniz of Fundacao Oswaldo Cruz (CPqGM-FIOCRUZ) Human Board, and all subjects provided written informed consent. For patients younger than 21 years, parents or guardians signed written informed consent followed by the child’s agreement. The study was developed in accordance with the Declaration of Helsinki of 1975 as revised in 2000. Clinical information was collected from the patients’ charts and their physicians.
Hemoglobin profile and hematological data
The hemoglobin profile was confirmed by high-performance liquid chromatography (HPLC) using the VARIANT I (Bio-Rad, CA, USA) equipment, which uses the principle of ion exchange. Calibrators were evaluated before the samples, according to the manufacturer’s recommendations.
Molecular biology analysis
Genomic DNA was extracted from leukocytes in 200 µL of peripheral blood using the DNA Kit (Qiagen, Foster City, CA, USA) according to the manufacturer’s recommendations. The DNA concentration was measured spectrophotometrically using the Nanodrop ND-1000 (Isogen Life Science, Amsterdam, The Netherlands).
Molecular biology analysis was performed by polymerase chain reaction (PCR) amplification using specific oligonucleotides for study-specific gene regions.
The restriction fragment length polymorphism (PCR-RFLP) technique was used to identify gene polymorphisms associated with the DIIIa and DAR alleles (Table 1). All samples containing the HincII restriction enzyme site had the 667C>G (exon 5) mutation in the RHD gene and were subjected to a second PCR reaction to search for the 455A>C (exon 3) and the 1025T>C (exon 7) mutations. The DNA fragment amplified by PCR was digested with the BanI restriction enzyme, to identify the 455A> C mutation specific for the DIIIa allele, and by the HphI restriction enzyme, to identify the 1025T>C mutation specific for the DAR allele [8, 9].
 Click to view | Table 1. Synthetic Oligonucleotides Used to Amplify Exons 3, 5, 7 and 8 of the RHD and HBB Genes |
The identification of the DAU allele was performed by PCR-SSP using the primers dau1b and daub to amplify the mutant allele located in exon 8 of the RHD gene [10]. The beta globin (HBB) gene was amplified as an internal control (Table 1) [11].
Statistical analysis
The analysis of the probability distribution of the variables was performed using the Kolmogorov-Smirnov test together with ANOVA parametric tests or non-parametric Kruskal-Wallis tests. The ANOVA test was used to analyze the distribution of means of quantitative or numerical variables with Gaussian distribution within categories. Moreover, to determine whether there was likely to be a difference between the mean values, we conducted multiple comparisons of means by the Bonferroni test (or post hoc). The non-parametric Kruskal-Wallis test was used for non-Gaussian distribution.
Analyses of qualitative or categorical variables with three or more groups were performed by non-parametric Chi-squared tests, properly corrected by Mantel-Haenszel and Yates tests. When values less than 4 were obtained, a Fisher’s exact test was used, and 95% confidence intervals (CIs) and prevalence ratios (PRs) were calculated for these variables.
The Mann-Whitney test and the independent t-test were used to analyze two quantitative variables by comparing two groups of values within a variable, taking into account their distribution.
The data analysis was performed using EPIInfo 6.04 (CDC, Atlanta, GA), the Statistics Data Analysis (STATA) SE 10 (StataCorp, TX, USA) and GraphPad Prism 5.0. A P value of less than 0.05 was considered statistically significant.
| Results | ▴Top |
Patient characteristics
We investigated 241 SCD patients with an average ± standard deviation (SD) age of 17.3 ± 10.9 years, of whom 134 (49.8%) were female. All of these subjects had hemoglobin patterns compatible with SCD, 171 (71%) with HbSS and 56 (29%) with SC disease (HbSC). A healthy control group was included in the study and was composed of 220 individuals with an average age of 36 ± 9.2 years, of whom 98 (44.2%) were female. All of the individuals in the healthy group had normal hemoglobin patterns.
RHD gene mutation frequencies
Tables 2 and 3 show the 667C>G (exon 5) mutation frequencies among the SCD and HbSS patients. The presence of the D partial phenotype was confirmed by searching for the DIIIa allele associated with the 455A>C mutation (exon 3), the DAR allele associated with the 1025T>C mutation (exon 7) and the DAU allele associated with the 1136C>T mutation in the RHD gene. All results were compared with the frequency of the same allele among healthy individuals from the control group.
 Click to view | Table 2. RHD Gene Mutation Frequencies Among the SCD Patients and Healthy Individuals |
 Click to view | Table 3. Distributions of DIIIa, DAR and DAU Alleles Frequencies in the RHD Gene Among Individuals With HbSS and Healthy Individuals |
Of 241 SCD patients selected, 193 were analyzed for the 667T>G (exon 5) mutation and 90.1% (174/193) had the wild-type allele, while 9.9% (19/193) had the mutant allele. Of these 19 SCD patients with the 667T>G mutant allele, 78.9% (15/19) were heterozygous, and 21.1% (4/19) were hemizygotes with a negative D allele (D667T>G/D-). The results from the group of healthy individuals showed that 95.9% (211/220) had the wild-type allele for the 667T>G mutation and 4.1% (9/220) had the mutant allele. Of these, 66.7% (6/9) were heterozygous, and 33.3% (3/9) were hemizygotes with a negative D allele. The 455 A>C mutation, which characterizes the DIIIa allele, was found in 2.1% (4/193) of SCD patients. Of these, 75% (3/4) were heterozygous, and 25% (1/4) were hemizygous (DIIIa /D-). Interestingly, the 455A>C mutation was not found among the group of healthy individuals. The 667T>G and 1136C>T (DAU) mutations were associated with DIIIa/DAR or 1136C>T (DAU) alleles that were not in Hardy-Weinberg equilibrium in either the SCD group or the HbSS group.
No difference was observed when the allele frequencies of SCD and HbSS patients groups were compared for 667T>G, 1136C>T (DAU), and the associated DIIIa/DAR or 1136C>T (DAU).
Distribution of DIIIa, DAR and DAU alleles
Table 4 shows the distribution of the DIIIa, DAR and DAU alleles in the RHD gene among the SCD, HbSS and HbSC disease patients.
 Click to view | Table 4. Distributions of the DIIIa, DAR and DAU Alleles in SCD and Healthy Groups |
Distribution of partial D genotype frequencies
Table 5 shows the genotype distribution of the DIIIa, DAR and DAU alleles of the RHD gene in the SCD, HbSS, and HbSC groups and in the group of healthy individuals. Patients with the D genotype associated with hemizygosity (D partial/D negative) or homozygosity or compound heterozygosity (D partial/D partial) were subject to alloimmunization. From a total of six patients, one (16.6%) had the genotype DIIIa/D-, three (50%) had the genotype DAR/D-, one (16.6%) had the genotype DIIIa/DAU, and one (16.6%) had the genotype DAR/DAU. Three (1.4%) of the healthy individuals were hemizygous and had the genotype DAR/D-.
 Click to view | Table 5. Distribution of Partial D Genotype Frequencies in the SCD, HbSS and Healthy Groups |
Distribution of the “C”, “c”, “E”, and “e” antigens of the Rh system and the DIIIa, DAR and DAU alleles of the RHD gene
Phenotypic data for the antigens “C”, “c”, “E” and “e” of the Rh system were investigated in the medical records of the patients. This information was available for 75 SCD and 60 HbSS patients. The most common phenotype found in the SCD group and the HbSS group was “ccee”, with 37.3% (28/75) and 36.7% (22/60) of patients demonstrating this phenotype, respectively. The second most common phenotype was “Ccee”, with 29.3% (22/75) in the SCD group and 30.0% (18/60) in the HbSS group.
In analysis of the frequencies of the phenotypes of the Rh system antigens in 36 patients with SCD who had a mutation in the RHD gene, the phenotype “ccee” was the most frequent at 56.8% (21/37), followed by “Ccee” at 18.9% (7/37) and “ccEe” at 18.9% (7/37) (Table 6).
 Click to view | Table 6. Distribution of the “C”, “c”, “E”, and “e” Antigens of the Rh System and the DIIIa, DAR and DAU Alleles of the Gene RHD Among Sickle Cell Disease Patients |
The distribution of the Rh system phenotypes among the SCD patients showed that the incidence of 667T>G and 1136C>T mutations with the R1R phenotype was more frequent among carriers of the wild-type allele, representing 36.4% (8/22) of the patients, followed by R0r with 22.7% (5/22). Regarding the presence of the 667T>G mutation and the 1136C>T mutation, the phenotype R0r was the most frequent, corresponding to 56.8% (21/37) of the patients, followed by the phenotype R1R with 18.9% (7/37).
The mutations 667T>G and 1025T>C, which are related to the DAR allele, were associated with a high risk of hospitalization (Fig. 1).
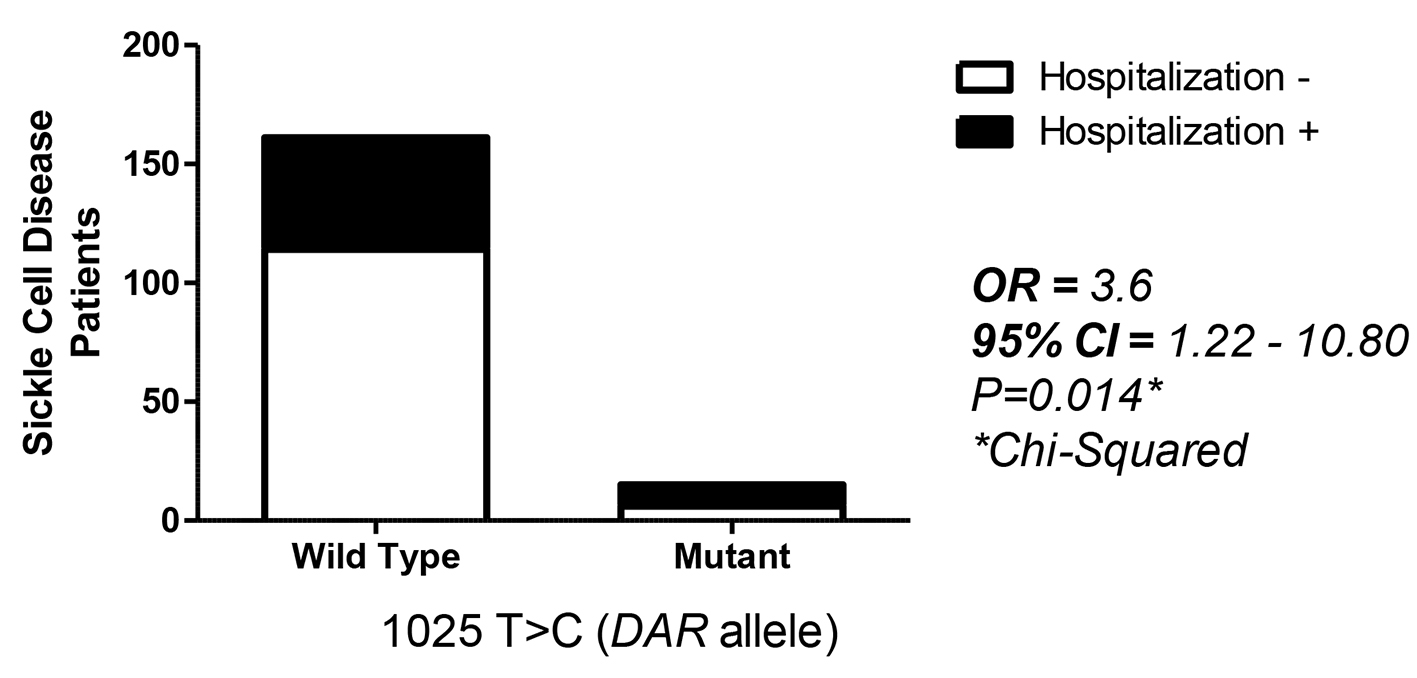 Click for large image | Figure 1. Distribution of 1025T>C, which is related to the DAR allele, and its association with a high risk of hospitalization. |
| Discussion | ▴Top |
The Rh system is extremely polymorphic and has important features related to ancestry, as changes described in this system are not homogeneously distributed among different ethnic groups [12]. Partial D antigens are rare in the Caucasian population but have high frequencies among individuals from Africa or of African descent [2-4].
Partial D antigens are involved in anti-D alloimmunization because they are recognized as weakly D positive by the monoclonal and polyclonal antisera typically used in serology. Individuals with these antigens usually have been transfused with Rh-positive RBC [13].
In this study, we investigated the distributions of the DIIIa, DAR and DAU alleles among SCD individuals from the Brazilian state of Bahia after identifying the 667T>G, 455A>C, 1025T>C, and 1136 C>T mutations in the RHD gene. Individuals with SCD that participated in this study had the highest frequencies of the DIIIa, DAR and DAU alleles. Of 241 SCD patients, 25.7% were positive for at least one of the RHD gene mutations investigated. Our data corroborate previous reports that showed high frequencies of the partial D alleles among African people [3, 4, 14].
The 667C>G mutation is associated with both the DIIIa and DAR alleles and was therefore used as an indicator of either of these alleles. The 667C>G mutation was frequent among SCD patients, with a 2.5-fold higher probability for the presence of the DAR or DIIIa allele in SCD patients compared with healthy individuals. When the HbSS patient group was considered, the 667C>G mutation was found at a higher frequency, with a 2.9-fold higher probability of being found among patients compared with individuals from the healthy group. However, no differences were found between the allele frequencies in HbSC patients and HbSS patients, indicating that the alloimmunization risk is high even among HbSS patients. The observation of a higher allele frequency among SCD patients could represent a selection of patients with severe disease that attend the outpatient clinic and have a high risk of developing RhD antibodies, increasing the risk of alloimmunization.
In our work, 2.1% of SCD patients had the 667C>G and the 455A>C mutations, indicating the presence of the DIIIa allele. Only one patient was hemizygous for this allele. The DAU and DAR alleles were found in 22.3% and 7.8% of the patients, respectively. Castilho et al in 2005 reported the DIIIa allele frequency among Brazilian SCD patients from the city of Campinas, located in the South of Brazil, a region with a predominance of Caucasian individuals, where 9.2% of patients had the 667C>G mutation, comprising 3.1% of the genotype DIIIa/D wild type, and 6.1% had the genotype DIIIa/D-. While Castilho et al in 2005 reported that hemizygous individuals (DIIIa/D-) predominated, the most common genotype in our study was DIIIa/D normal [9]. However, the finding of a greater number of hemizygous individuals (DIIIa/D-) may suggest an increased risk of alloimmunization among SCD patients from Campinas.
The DIIIa allele was not found among individuals from the control group, and analysis of this allele among SCD, HbSS and HbSC patients and the healthy group did not show significant differences. Three of four patients with the 667C>G mutation were from the HbSS group, providing an estimate of the incidence of a partial D genotype DIIIa, of 2.2% of these patients. These findings corroborate the observation that a high frequency of this mutation is common in people of African origin and uncommon among individuals with a Caucasian genetic heritage [8].
We found the DAR allele in 7.8% of SCD patients and in 8.8% of HbSS patients, which is higher than reported in other studies. Hemker et al studied 326 individuals from South Africa and reported a frequency of 4.9% for the DAR allele, representing 1.5% of all homozygotes or hemizygotes [15]. In populations of Caucasian origin, however, the frequency of the DAR allele is lower than in populations of African origin, with reported frequencies lower than 0.1% [10]. The frequency of the DAR allele reported by Avent in 2009 was 0.0002%, while the frequency of the DAU allele was 0.0001% among 33,864 healthy individuals from a German population. In our study, the frequency of the DAR variant was 7.8% of the mutant alleles in SCD patients [16]. Castilho et al in 2005 described a frequency of 6.9% for this allele in SCD patients from Campinas [9].
A study conducted in the region of Cape Town, South Africa, demonstrated that approximately 11% of pregnant women had Rh-positive anti-D. Molecular biology analysis revealed that all of these women were positive for a D partial allele [6]. This variant is very common in African populations according to previous reports [17]. In 2009, Hustinx et al studied people of African descendant and found a frequency of 24.5% for the DAU allele identified by PCR-SSP [18]. In Brazil, the DAU allele has not been reported previously, and the high frequency of this allele among our SCD patients demonstrates the relevance of this work. In our study, the variant DAU was identified by a single codon mutation, 1136C>T [10]. Thus, the DAU allele, compared with DIIIa and DAR, was the most frequent in the SCD patients studied. These data reinforce the report that the DAU allele, not the DIIIa allele, is the most frequent RHD allele in the Brazilian population of Bahia, followed by the DAR allele. Castilho et al, in 2005, found 9/130 (6.9%) HbSS patients with the DAR allele, but only 5/130 (3.9%) with the DIIIa allele [9]. On the basis of the results obtained here, the DAU allele may be associated with a heterogeneous distribution of hemoglobin S (HbS) in Brazil, but this allele was not investigated in this study. In Northeast Brazil, mainly in Bahia, the population is a tri-racial mixture of Europeans (mostly Portuguese), Africans and indigenous people. The frequency of HbS in the state of Bahia is the highest in Brazil, varying from 4.5% to 14.7% in several population groups studied [7].
The distribution of the “C”, “c”, “E” and “e” antigens of the Rh system, according to positivity for the 667T>G and 1136C>T mutations, indicated that the phenotype R1R was the most common at 36.4% (8/22) of patients, followed by R0r with 22.7% (5/22). Among patients who were positive for any of the mutations, the most common phenotype was R0r at 56.8% (21/37), followed by R1R at 18.9% (7/37). Another study detected a DAR variant allele correlated with the “ce” phenotype in an Afro-Caribbean woman [19]. A group of European Caucasians individuals with the 1136C>T mutation exhibited the R0R2 phenotype more frequently than the Brazilian population [18].
For a similar Bahia population, D-typing strategies may be appropriate because of the variety of alleles presented by African clusters that are difficult to identify. In the case of SCD patients, this picture is even worse. In patients with anemia, the disease may be aggravated due to hemolysis caused by a transfusion reaction. This may decrease the effectiveness of transfusion therapy, generating free hemoglobin with the potential to damage kidneys and lungs, and increasing the level of reactive oxygen species, which can aggravate the disease pathogenesis [20]. These effects may partly account for the 3.6-fold increase in the risk for hospitalization in the SCD patients with the DAR allele in our study and indicate the need for improving transfusion therapy, for example, by genotyping patients for DAR, DIIIa and DAU, especially those who are candidates for chronic transfusion, to prevent alloimmunization [21].
Conclusions
Distributions of the variants of DIIIa, DAR, and DAU among SCD patients from the Brazilian state of Bahia differ from the frequencies observed in healthy individuals from the same population. The data presented here show that the DIIIa, DAR and DAU alleles are common in SCD and HbSS patients when analyzed separately, with a high frequency of the DAU allele; this is the first description of these high incidences in Brazil. The variant DAU was associated with the occurrence of anti-D alloimmunization among the SCD patients investigated. However, further studies should be performed to confirm the alloimmunization occurrence in two double hemizygous DAR/D- and DIIIa/D- patients who were submitted to several transfusion events, but had negative indirect Coombs.
Acknowledgments
This work was supported by grants from the Brazilian National Council of Research (CNPq) (3065427/2007-5 and 484457/2007-1) (M.S.G.); the Foundation of Research and Extension of Bahia (FAPESB) (1431040053063; 9073/2007 and 6234/2010) (M.S.G.); and MCD/CNPq/MS-SCTIE-DECIT (409800/2006-6) (M.S.G.). The sponsors of this study are public or nonprofit organizations that support science in general. They had no role in gathering, analyzing or interpreting the data.
Competing Interests
The authors declare no competing financial interests.
| References | ▴Top |
- Avent ND, Reid ME. The Rh blood group system: a review. Blood. 2000;95(2):375-387.
pubmed - Wagner FF, Flegel WA. RHD gene deletion occurred in the Rhesus box. Blood. 2000;95(12):3662-3668.
pubmed - Wagner FF, Flegel WA. Review: the molecular basis of the Rh blood group phenotypes. Immunohematology. 2004;20(1):23-36.
pubmed - Flegel WA, Wagner FF. Molecular biology of partial D and weak D: implications for blood bank practice. Clin Lab. 2002;48(1-2):53-59.
pubmed - Tippett P, Lomas-Francis C, Wallace M. The Rh antigen D: partial D antigens and associated low incidence antigens. Vox Sang. 1996;70(3):123-131.
doi pubmed - du Toit ED, Martell RW, Botha I, Kriel CJ. Anti-D antibodies in Rh-positive mothers. S Afr Med J. 1989;75(9):452.
pubmed - Azevedo ES, Alves AF, Da Silva MC, Souza MG, Muniz Dias Lima AM, Azevedo WC. Distribution of abnormal hemoglobins and glucose-6-phosphate dehydrogenase variants in 1200 school children of Bahia, Brazil. Am J Phys Anthropol. 1980;53(4):509-512.
doi pubmed - Westhoff CM, Vege S, Halter-Hipsky C, Whorley T, Hue-Roye K, Lomas-Francis C, Reid ME. DIIIa and DIII Type 5 are encoded by the same allele and are associated with altered RHCE*ce alleles: clinical implications. Transfusion. 2010;50(6):1303-1311.
doi pubmed - Castilho L, Rios M, Rodrigues A, Pellegrino J, Jr., Saad ST, Costa FF. High frequency of partial DIIIa and DAR alleles found in sickle cell disease patients suggests increased risk of alloimmunization to RhD. Transfus Med. 2005;15(1):49-55.
doi pubmed - Wagner FF, Ladewig B, Angert KS, Heymann GA, Eicher NI, Flegel WA. The DAU allele cluster of the RHD gene. Blood. 2002;100(1):306-311.
doi pubmed - Goncalves MS, Nechtman JF, Figueiredo MS, Kerbauy J, Arruda VR, Sonati MF, Saad SO, et al. Sickle cell disease in a Brazilian population from Sao Paulo: a study of the beta s haplotypes. Hum Hered. 1994;44(6):322-327.
doi pubmed - Cartron JP. RH blood group system and molecular basis of Rh-deficiency. Baillieres Best Pract Res Clin Haematol. 1999;12(4):655-689.
doi pubmed - Daniels G. Human Blood Groups. 2nd ed. Blackwell, Oxford, 2002.
- Wagner FF, Eicher NI, Jorgensen JR, Lonicer CB, Flegel WA. DNB: a partial D with anti-D frequent in Central Europe. Blood. 2002;100(6):2253-2256.
doi pubmed - Hemker MB, Ligthart PC, Berger L, van Rhenen DJ, van der Schoot CE, Wijk PA. DAR, a new RhD variant involving exons 4, 5, and 7, often in linkage with ceAR, a new Rhce variant frequently found in African blacks. Blood. 1999;94(12):4337-4342.
pubmed - Avent ND. Large-scale blood group genotyping: clinical implications. Br J Haematol. 2009;144(1):3-13.
doi pubmed - Wagner FF, Moulds JM, Tounkara A, Kouriba B, Flegel WA. RHD allele distribution in Africans of Mali. BMC Genet. 2003;4:14.
doi pubmed - Hustinx H, Poole J, Bugert P, Gowland P, Still F, Fontana S, Scharberg EA, et al. Molecular basis of the Rh antigen RH48 (JAL). Vox Sang. 2009;96(3):234-239.
doi pubmed - Silvy M, Chapel-Fernandes S, Callebaut I, Beley S, Durousseau C, Simon S, Lauroua P, et al. Characterization of novel RHD alleles: relationship between phenotype, genotype, and trimeric architecture. Transfusion. 2012;52(9):2020-2029.
doi pubmed - Noizat-Pirenne F. Relevance of alloimmunization in haemolytic transfusion reaction in sickle cell disease. Transfus Clin Biol. 2012;19(3):132-138.
doi pubmed - Miller ST, Kim HY, Weiner DL, Wager CG, Gallagher D, Styles LA, Dampier CD, et al. Red blood cell alloimmunization in sickle cell disease: prevalence in 2010. Transfusion. 2013;53(4):704-709.
doi
This article is distributed under the terms of the Creative Commons Attribution Non-Commercial 4.0 International License, which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Journal of Hematology is published by Elmer Press Inc.